MUGO: Multi-Head Genomic Optimization
🛑 IMPORTANT FOR REVIEWERS
This repository is anonymized for KDD 2026 double-blind review. You can access the full source code via the Anonymous GitHub Repository.
MUGO is a differentiable combinatorial optimization framework designed for discovering causal variants in the non-coding genome. By leveraging Gumbel-Softmax relaxation and Straight-Through Estimators (STE), MUGO enables end-to-end gradient-based optimization on discrete DNA sequences.
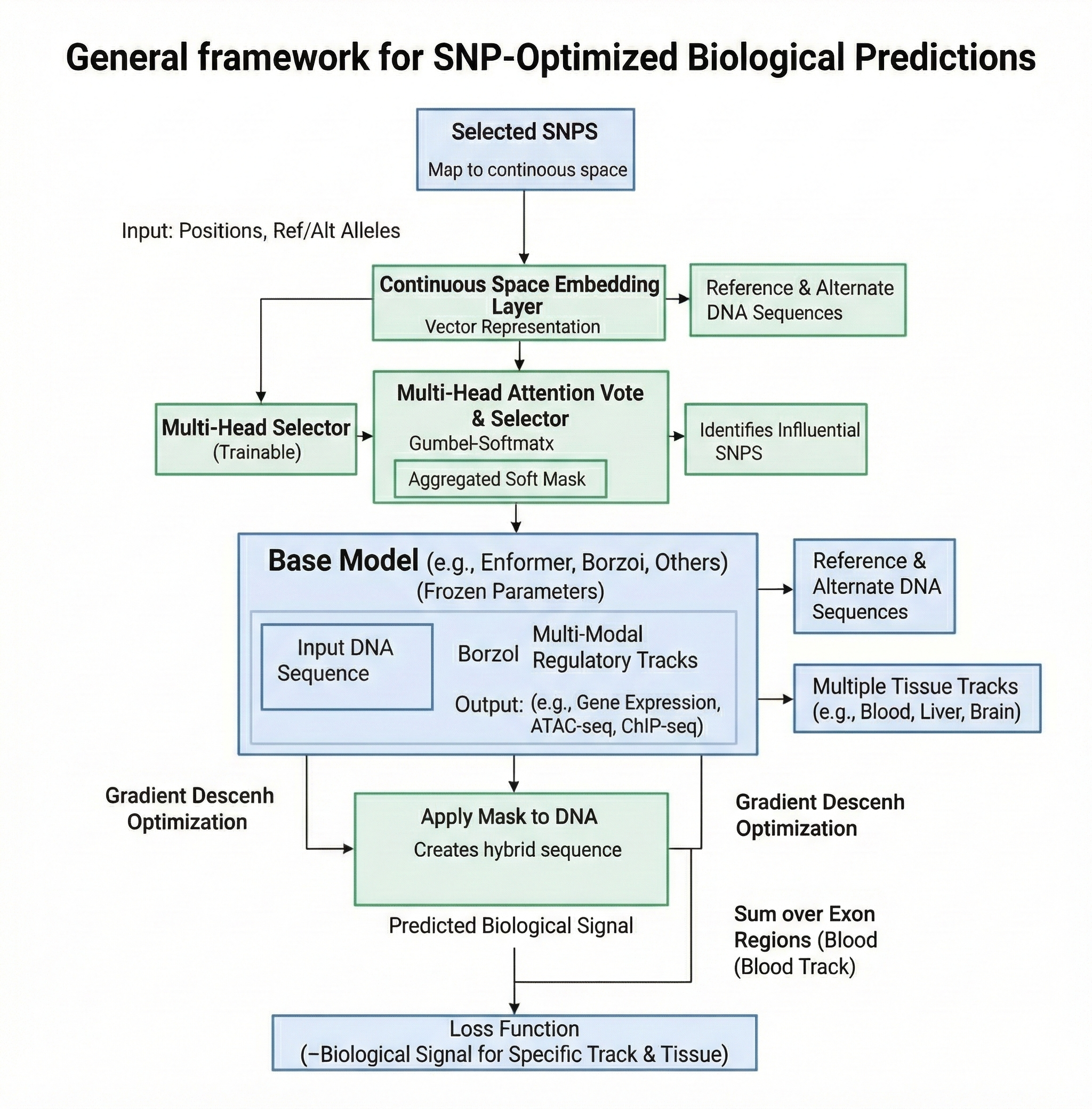
Key Features
- 🧬 Model-Agnostic: Compatible with Borzoi, Enformer, HyenaDNA, and other PyTorch-based genomic models.
- 🎯 Multi-Modal Objectives: Optimize for Gene Expression, Chromatin Accessibility (ATAC), or TF Binding.
- 📉 Variance Reduction: Built-in Multi-Head Consensus strategy to filter stochastic noise.
- 🚀 Production Ready: Easy-to-use Python API for high-performance computing.
Citation
If you use MUGO in your research, please cite our KDD 2026 paper (under review).